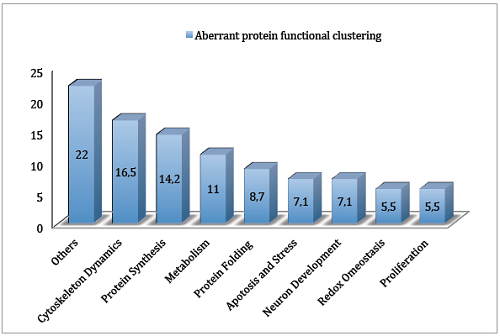
Differential protein expression in normal and trisomic cell lines from murine cerebral cortex measured by itraq and Mass Spectrometry
Abstract
Full Text:
PDFReferences
Antonarakis SE, 10 years of genomics, chromosome 21, and Down syndrome. Genomics, 1998, 51, 1-16.
Penrose LS, The relative effects of paternal and maternal age in mongolism. J Genet., 1933, 27, 219-224.
Penrose LS, Mongolian idiocy (mongolism) and maternal age. Ann N Y Acad Sci., 1954, 57:494-502.
Epstein CJ, Down syndrome. The metabolic and molecular bases of inherited disease, B. A. Scriver CR, Sly W, Valle D. New York, McGraw Hill, Inc., 1995, 1, 749-794.
Raz N, Torres IJ, Briggs SD, Spencer WD, Thornton AE, Loken WJ, Gunning FM, McQuain JD, Driesen NR, Acker JD, Selective neuroanatomic abnormalities in Down’s syndrome and their cognitive correlates: evidence from MRI morphometry, Neurology, 1995, 45, 356-366.
Schmidt-Sidor B, Wisniewski KE, Shepard TH, Sersen EA, Brain growth in Down syndrome subjects 15 to 22 weeks of gestational age and birth to 60 months, Clin Neuropathol., 1990, 9, 181-190.
Golden JA, Hyman BT, Development of the superior temporal neocortex is anomalous in trisomy 21, J Neuropathol Exp Neurol., 1994, 53, 513-520.
Wisniewski KE, Down syndrome children often have brain with maturation delay, retardation of growth, and cortical dysgenesis, Am J Med Genet Suppl., 1990, 7, 274-281.
Ãbrahám H, Vincze A, Veszprémi B, Kravják A, Gömöri É, Kovács GG, Seress L. Impaired myelination of the human hippocampal formation in Down syndrome, Int J Dev Neurosci., 2012, 30, 147-58.
Lögdberg B, Brun A, Prefrontal neocortical disturbances in mental retardation. J Intellect Disabil Res., 1993, 37, 459-468.
Ross MH, Galaburda AM, Kemper TL, Down’s syndrome: Is there a decreased population of neurons?, Neurology, 1984, 34, 909-916.
Weitzdoerfer R, Dierssen M, Fountoulakis M, Lubec G, Fetal life in Down syndrome starts with normal neuronal density but impaired dendritic spines and synaptosomal structure. J Neural Transm Suppl., 2001, 61, 59-70.
Epstein CJ, Cox DR, Epstein LB, Mouse trisomy 16: an animal model of human trisomy 21 (Down syndrome), Ann N Y Acad Sci., 1985, 450, 157-168.
Galdzicki Z, Siarey RJ, Understanding mental retardation in Down's syndrome using trisomy 16 mouse models Genes, Brain and Behavior, 2003, 2, 167–178.
Allen DD, Martin J, Arrigada C, Cardenas AM, Rapoport SI, Caviedes R, Caviedes P, Impaired cholinergic function in cell Unes derived from the cerebral cortex of normal and trisomy 16 mice, Eur J Neurosci., 2000, 12, 3259-3264.
Cárdenas AM, RodrÃguez MP, Cortés MP, Alvarez RM, Wei W, Rapoport SI, Shimahara T, Caviedes R, Caviedes P, Calcium signals in cell lines derived from the cerebral cortex of normal and trisomy 16 mice, Neuroreport, 1999, 10, 363-369.
Saud K, Arrigada C, Cardenas AM, Shimahara T, Allen DD, Caviedes R, Caviedes P, Neuronal dysfunction in Down syndrome: Contribution of neuronal models in cell culture, J Physiol Paris, 2006, 99, 201-210.
Paula Lima AC, Arriagada C, Toro R, Cárdenas AM, Caviedes R, Ferreira ST, Caviedes, P, Small-molecule aggregation inhibitors reduce excess amyloid in a trisomy 16 mouse cortical cell line, Biol Res., 2008, 41, 29-36.
Arrigada C, Astorga C, Atwater I, Rojas E, Mears D, Caviedes R R, Endosomal abnormalities related to amyloid precursor protein in cholesterol treated cerebral cortex neuronal cells derived from trisomy 16 mice, an animal model of Down syndrome. Neurosci Lett., 2007, 423, 173-177.
Arriagada C, Bustamante M, Atwater I, Rojas E, Caviedes R, Caviedes P, Apoptosis is directly related to intracellular amyloid accumulation in a cell line derived from the cerebral cortex of a trisomy 16 mouse, an animal model of Down syndrome. Neurosci Lett., 2010, 470, 81-5.
Bacchus C, Sterz H, Buselmaier W, Sahai S, Winking H, Genesis and systematization of cardiovascular anomalies and analysis of skeletal malformations in murine trisomy 16 and 19: two animal models for human trisomies. Hum Genet, 1987, 77, 12-22.
Webb S, Anderson RH, Brown NA, Endocardial cushion development and heart loop architecture in the trisomy 16 mouse. Dev Dyn., 1996, 206, 301-309.
Grausz H, Richtsmeier JT, Oster-Granite ML, Morphogenesis of the brain and craniofacial complex in trisomy 16 mice, In: The morphogenesis of Down syndrome, (Epstein CJ, ed), New York: Wiley-Liss, Inc., 1991, pp.169-188.
Ewart JL, Auerbach R, Defects in thymocyte differentiation and thymocyte-stromal interactions in the trisomy 16 mouse. Dev Immunol., 1992, 2, 215-226.
Gjertson C, Sturm KS, Berger CN, Hematopoietic deficiencies and core binding factor expression in murine ts16, an animal model for Down syndrome. Clin Immunol., 1999, 91, 50-60
Haydar TF, Blue ME, Molliver ME, Kreuger BK, Yarowsky PJ, Consequences of trisomy 16 for mouse brain development: corticogenesis in a model of Down syndrome. J Neurosci., 1996, 16, 6175-6182.
Orozco CB, Smith SA, Epstein CJ, Rapoport SI, Electrophysiological properties of cultured dorsal root ganglion and spinal cord neurons of normal and trisomy 16 fetal mice. Brain Res., 1986, 429, 111-122.
Taniguchi Y, Choi PJ, Li GW, Chen H, Babu M, Hearn J, Emili A, Xie XS, Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science, 2010, 329, 533-538.
Mao R, Zielke CL, Zielke HR, Pevsner J. Global up-regulation of chromosome 21 gene expression in the developing Down syndrome brain. Genomics, 2003, 81, 457-467.
Amano K, Sago H, Uchikawa C, Suzuki T, Kotliarova SE, Nukina N, Epstein CJ, Yamakawa K , Dosage-dependent over-expression of genes in the trisomic region of Ts1Cje mouse model for Down syndrome, Hum Mol Genet, 2004, 13, 1333-1340.
Kahlem P, Sultan M, Herwig R, Steinfath M, Balzereit D, Eppens B, Saran NG, Pletcher MT, South ST, Stetten G, Lehrach H, Reeves RH, Yasps ML, Transcript level alterations reflect gene dosage effects across multiple tissues in a mouse model of Down syndrome, Genome Res., 2004, 14, 1258-1267.
Lyle R, Gehrig C, Neergaard-Henrichsen C, Deutsch S, Antonarakis SE, Gene expression from the aneuploid chromosome in a trisomy mouse model of Down syndrome, Genome Res., 2004, 14, 1268-1274.
Epstein CJ, Down's syndrome: critical genes in a critical region. Nature, 2006, 441, 582–583.
Aït Yahya-Graison E, Aubert J, Dauphinot L, Rivals I, Prieur M, Golfier G, Rossier J, Personnaz L, Creau N, Bléhaut H, Robin S, Delabar JM, Potier MC, Classification of human chromosome 21 gene-expression variations in Down syndrome: impact on disease phenotypes, Am J Hum Genet., 2007, 81, 475-91.
Prandini P, Deutsch S, Lyle R, Gagnebin M, Delucinge Vivier C, Delorenzi M, Gehrig C, Descombes P, Sherman S, Dagna Bricarelli F, Baldo C, Novelli A, Dallapiccola B, Antonarakis SE, Natural gene-expression variation in Down syndrome modulates the outcome of gene-dosage imbalance, Am J Hum Genet., 2007, 81, 252-63.
Engidawork E, Gulesserian T, Fountoulakis M, Lubec G, Aberrant protein expression in cerebral cortex of fetus with Down syndrome, Neuroscience, 2003, 122, 145-54.
Becker L, Mito T, Takashima S, Onodera K, Growth and development of the brain in Down syndrome, Prog Clin Biol Res., 1991, 373, 133-152.
Cotman CW, Monaghan DT, Ganong AH, Excitatory amino acid neurotransmission: NMDA receptors and Hebb-type synaptic plasticity, Annu Rev Neurosci., 1988, 11, 61– 80.
Palaiologos G, Hertz L, Schousboe A, Evidence that aspartate aminotransferase activity and ketodicarboxylate carrier function are essential for biosynthesis of transmitter glutamate, J Neurochem., 1988, 51, 317–320.
Bao X, Pal R, Hascup KN, Wang Y, Wang WT, Xu W, Hui D, Agbas A, Wang X, Michaelis ML, Choi IY, Belousov AB, Gerhardt GA, Michaelis EK, Transgenic expression of Glud1 (glutamate dehydrogenase 1) in neurons: in vivo model of enhanced glutamate release, altered synaptic plasticity, and selective neuronal vulnerability, J Neurosci., 2009, 29, 13929-13944.
Siarey RJ, Stoll J, Rapoport SI, Galdzicki Z, Altered long-term potentiation in the young and old Ts65Dn mouse, a model for Down Syndrome, Neuropharmacology, 1997, 36, 1549-1554.
Bliss TV, Collingridge GL, A synaptic model of memory: long-term potentiation in the hippocampus, Nature, 1993, 361, 31–39.
Cooke SF, Bliss TV, Plasticity in the human central nervous system, Brain, 2006, 129, 1659–73.
Lee AS, Mammalian stress response: induction of the glucose-regulated protein family. Curr Opin Cell Biol., 1992, 4, 267–273.
Zhao L, Rosales C, Seburn K, Ron D, Ackerman SL, Alteration of the unfolded protein response modifies neurodegeneration in a mouse model of Marinesco-Sjogren syndrome, Hum Mol Genet., 2010, 19, 25-35.
Brodsky JL, Werner ED, Dubas ME, Goeckeler JL, Kruse KB, and McCraken AA, The requirement for molecular chaperones during endoplasmic reticulum-associated protein degradation demonstrates that protein export and import are mechanistically distinct, J Biol Chem., 1999, 274, 3453–3460.
Gething, MJ, Role and regulation of the ER chaperone BiP, Semin Cell Dev Biol., 1999, 10, 465–472.
Forman MS, Lee VM, Trojanowski JQ, ‘Unfolding’ pathways in neurodegenerative disease, Trends Neurosci., 2003, 26, 407–410.
Kaufman RJ, Stress signaling from the lumen of the endoplasmic reticulum: coordination of gene transcriptional and translational controls. Genes Dev., 1999, 13, 1211–1233.
Kaufman RJ, Orchestrating the unfolded protein response in health and disease, J Clin Invest., 2002, 110, 1389-1398.
Salminen A, Kauppinen A, Suuronen T, Kaarniranta K, Ojala J, ER stress in Alzheimer's disease: a novel neuronal trigger for inflammation and Alzheimer's pathology, J Neuroinflammation, 2009, 6, 41.
Yao Y, Zhou Y, and Wang C, Both the isomerase and chaperone activities of protein disulfide isomerase are required for the reactivation of reduced and denatured acidic phospholipase A2, EMBO J., 1997, 16: 651–658.
Chen B, Piel WH, Gui L, Bruford E, Monteiro A. The HSP90 family of genes in the human genome: insights into their divergence and evolution, Genomics. 2005, 86, 627-37.
Wakatsuki T, Hatayama T, Characteristic expression of 105-kDa heat shock protein (HSP105) in various tissues of nonstressed and heat-stressed rats, Biol Pharm Bull., 1998, 21, 905–910.
Yamagishi N, Nishihori H, Ishihara K, Ohtsuka K, Hatayama T, Modulation of the chaperone activities of Hsc70/Hsp40 by Hsp105a and Hsp105b, Biochem Biophys Res Commun., 2000, 272, 850– 855.
Dragovic Z, Broadleu SA, Shomura Y, Bracher A, Hartl FU, Molecular chaperones of the Hsp110 family act as nucleotide exchange factors of Hsp70s, EMBO J., 2006, 25, 2519-2528.
Hatayama T, Yamagishi N, Minobe E, Sakai K, Role of hsp105 in protection against stress-induced apoptosis in neuronal PC12 Cells, Biochem Biophys Res Commun., 2001, 288:528–534.
Saito Y, Yamagishi N, Ishihara K, Hatayama T, Identification of alpha-tubulin as an hsp105alpha-binding protein by the yeast two-hybrid system, Exp Cell Res., 2003, 286:233-240.
Satoh H, Shibata H, Nakano Y, Kitaura Y, Maki M, ALG-2 interacts with the amino-terminal domain of annexin XI in a Ca(2+)-dependent manner, Biochem Biophys Res Commun., 2002, 291, 1166-1172.
Kuhn DE, Nuovo GJ, Terry AV Jr, Martin MM, Malana GE, Sansom SE, Pleister AP, Beck WD, Head E, Feldman DS, Elton TS. Chromosome 21-derived microRNAs provide an etiological basis for aberrant protein expression in human Down syndrome brains, J Biol Chem., 2010, 285, 1529-43.
Kaufmann WE Mental retardation and learning disabilities: A neuropathologic differentiation, In: Developmental disabilities in infancy and childhood, (Capute AJ, Accardo PJ, eds), Baltimore, MD: Paul H. Brookes Publishing, 1996, 2, 49–70.
Pollard TD, Borisy GG, Cellular motility driven by assembly and disassembly of actin filaments, Cell., 2003, 112, 453–465.
Ramesh V, Merlin and the ERM proteins in Schwann cells, neurons and growth cones, Nat Rev Neurosci., 2004, 5, 462–470.
Haas MA, Vickers JC, Dickson TC, Rho kinase activates ezrin-radixin-moesin (ERM) proteins and mediates their function in cortical neuron growth, morphology and motility in vitro, J Neurosci Res., 2007, 85, 34-46.
Giretti MS, Fu XD, De Rosa G, Sarotto I, Baldacci C, Garibaldi S, Mannella P, Biglia N, Sismondi P, Genazzani AR, Simoncini T, Extra-nuclear signalling of estrogen receptor to breast cancer cytoskeletal remodelling, migration and invasion. PLoS ONE, 2008, 3(5), e2238.
Sanchez AM, Flamini MI, Fu XD, Mannella P, Giretti MS, Goglia L, Genazzani AR, Simoncini T, Rapid signaling of estrogen to WAVE1 and moesin controls neuronal spine formation via the actin cytoskeleton. Mol Endocrinol., 2009, 23, 1193-1202.
Hirokawa N, Kinesin and dynein superfamily proteins and the mechanism of organelle transport. Science, 1998, 279, 519-526.
Schliwa M, Woehlke G, Molecular motors, Nature, 2003, 422:759-765.
Vale RD, The molecular motor toolbox for intracellular transport. Cell., 2003, 112, 467-480.
Vallee RB, Williams JC, Varma D, Barnhart LE, Dynein: an ancient motor protein involved in multiple modes of transport. J Neurobiol., 2004, 58, 189-200.
Heerssen HM, Pazyra MF, Segal RA, Dynein motors transport activated Trks to promote survival of target-dependent neurons. Nat Neurosci., 2004, 7, 596–604.
Ibañez-Tallon I, Pagenstecher A, Fliegauf M, Olbrich H, Kispert A, Ketelsen UP, North A, Heintz N, Omran H, Dysfunction of axonemal dynein heavy chain Mdnah5 inhibits ependymal flow and reveals a novel mechanism for hydrocephalus formation. Hum Mol Genet., 2004, 13, 2133-2141.
Forcelini CM, Mallmann AB, Crusius PS, Seibert CA, Crusius MU, Zandoná DI, Carazzo C, Crusius CU, Goellner E, Ragnini J, Manzato LB, Winkelmann G, Lima AV, Bauermann MG, Down syndrome with congenital hydrocephalus: case report. Arq Neuro-psiquiatr., 2006, 64, 869-871.
Jayaraman A, Ballweg GP, Donnenfeld H, Chusid JG, Hydrocephalus in Down's syndrome. Childs Brain, 1976, 2, 202-207.
Yu T, Li Z, Jia Z, Clapcote SJ, Liu C, Li S, Asrar S, Pao A, Chen R, Fan N, Carattini-Rivera S, Bechard AR, Spring S, Henkelman RM, Stoica G, Matsui S, Nowak NJ, Roder JC, Chen C, Bradley A, Yu YE, A mouse model of Down syndrome trisomic for all human chromosome 21 syntenic regions. Hum Mol Genet., 2010, 19, 2780-2791.
Malamud N, Neuropathology of organic brain syndromes associated with aging. In: Aging Brain (Gaitz CM ed), New York: Plenum Press, 1972, pp 63-87.
Kimura N, Inoue M, Okabayashi S, Ono F, Negishi T, Dynein dysfunction induces endocytic pathology accompanied by an increase in Rab GTPases: a potential mechanism underlying age-dependent endocytic dysfunction. J Biol Chem., 2009, 284, 31291-31302.
Borer RA, Lehner CF, Eppenberger HM, Nigg EA, Major nucleolar proteins shuttle between nucleus and cytoplasm. Cell., 1989, 56, 379–390.
Yu Y, Maggi LB Jr., Brady SN, Apicelli AJ, Dai MS, Lu H, Nucleophosmin is essential for ribosomal protein L5 nuclear export. Mol Cell Biol., 2006, 26, 3798-3809.
Herrera JE, Savkur R, Olson MO, The ribonuclease activity of nucleolar protein B23. Nucleic Acids Res., 1995, 23, 3974-3979.
Wang D, Baumann A, Szebeni A, Olson MO, The nucleic acid binding activity of nucleolar protein B23.1 resides in its carboxyl- terminal end. J Biol Chem., 1994, 269, 30994-30998.
Szebeni A, Olson MO, Nucleolar protein B23 has molecular chaperone activities. Protein Sci., 1999, 8, 905-912.
Grisendi S, Bernardi R, Rossi M, Cheng K, Khandker L, Manova K, Pandolfi PP, Role of nucleophosmin in embryonic development and tumorigenesis, Nature, 2005, 437, 147-153.
Falini B, Mecucci C, Tiacci E, Alcalay M, Rosati R, Pasqualucci L, La Starza R, Diverio D, Colombo E, Santucci A, Bigerna B, Pacini R, Pucciarini A, Liso A, Vignetti M, Fazi P, Meani N, Pettirossi V, Saglio G, Mandelli F, Lo-Coco F, Pelicci PG, Martelli MF, Cytoplasmic nucleophosmin in acute myelogenous leukemia with a normal karyotype. N Engl J Med., 2005, 352, 254–266.
Xavier AC, Taub JW, Acute leukemia in children with Down syndrome. Haematologica, 2010, 95, 1043-1045.
Stewart CF, Treating children with acute lymphoblastic leukemia and Down syndrome: pharmacokinetics provides insight into vincristine therapy. Pediatr Blood Cancer, 2009, 52, 1-2.
Lange BJ, Kobrinsky N, Barnard DR, Arthur DC, Buckley JD, Howells WB, Gold S, Sanders J, Neudorf S, Smith FO, Woods WG, Distinctive demography, biology, and outcome of acute myeloid leukemia and myelodysplastic syndrome in children with Down syndrome: Children's Cancer Group Studies 2861 and 2891. Blood, 1998, 91, 608-615.
Shilov IV, Seymour SL, Patel AA, Loboda A, Tang WH, Keating SP, Hunter CL, Nuwaysir LM, Schaeffer DA,The Paragon Algorithm, a next generation search engine that uses sequence temperature values and feature probabilities to identify peptides from tandem mass spectra. Mol Cell Proteomics, 2007, 6, 1638-55.
Jain E, Bairoch A, Duvaud S, Phan I, Redaschi N, Suzek BE, Martin MJ, McGarvey P., Gasteiger E, Infrastructure for the life sciences: design and implementation of the UniProt website BMC Bioinformatics, 2009, 10, 136.
Blake JA, Bult CJ, Kadin JA, Richardson JE, Eppig JT and the Mouse Genome Database Group, The Mouse Genome Database (MGD): premier model organism resource for mammalian genomics and genetics. Nucleic Acids Res., 2011, 39 (suppl 1): D842-D84.
DOI: http://dx.doi.org/10.13171/mjc.2.2.2012.12.12.06
Refbacks
- There are currently no refbacks.
Copyright (c) 2015 Mediterranean Journal of Chemistry
